Cellular RNA folding is a highly dynamic process, occurring as RNA is synthesized. Called cotranscriptional folding, this process happens each time an RNA is made in the cell, and it is thought to influence myriad gene-expression processes as well as how RNAs attain their functional states. Despite its importance, we have never had a clear picture of the mechanisms by which RNAs fold cotranscriptionally. This gap limits understanding of the most basic aspects of RNA biology, such as how RNAs called riboswitches dynamically alter their structures in response to ligand binding to act as cellular sensors, how competing RNA folds can inhibit or enhance RNA splicing, and how nascent RNA folds drive specific patterns of RNA modification.
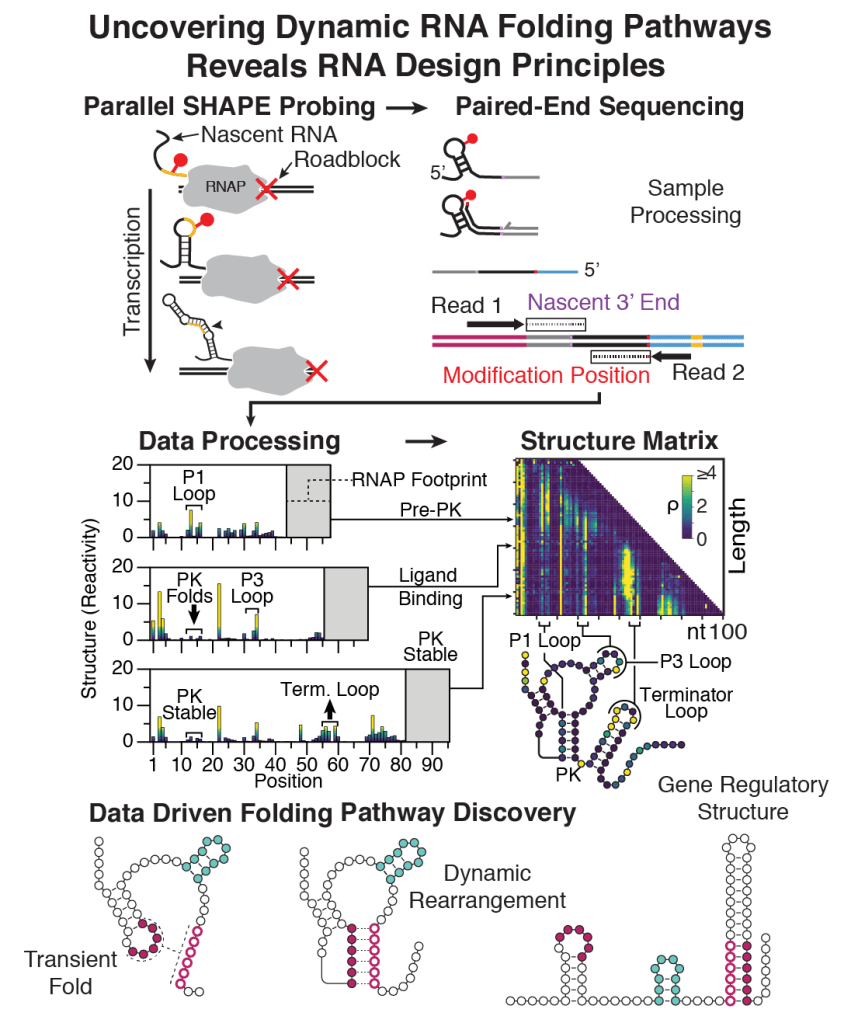
To address this challenge, we invented cotranscriptional SHAPE-Seq to reveal RNA cotranscriptional folding pathways at the highest structural resolution yet attained. We have used this technique to uncover the first nucleotide-resolution information about the folding pathways of two starkly different riboswitch sensor RNAs and have discovered that these pathways may share strikingly similar features. In ongoing work, we are developing new computational algorithms that can model entire cotranscriptional RNA folding pathways using SHAPE-Seq data, which is showing that the mechanistic similarities of riboswitches also underlie the folding pathway of one of the most ancient non-coding RNAs known, the signal recognition particle (SRP) RNA. We are currently very interested in learning how transcription dynamics impacts nascent RNA folding, how cellular transcription factors impact nascent RNA folding, and what are the common RNA folding mechanisms that allows RNAs to traverse kinetic traps during cotranscriptional folding.
Featured Publications:
1. Computationally reconstructing cotranscriptional folding pathways from experimental data reveals rearrangement of non-native folding intermediates.
A. M Yu#, P. M. Gasper#, E. J. Strobel, K. E. Watters, A. A. Chen*, J. B. Lucks*. (2018). # = Equal contribution
Links: BioRxiv
2. DUETT quantitatively identifies known and novel events in nascent RNA structural dynamics from chemical probing data.
A. Y Xu, A. M Yu, J. B. Lucks, N. Bagheri.* Bioinformatics. (2019).
Links: Journal, PDF
3. A ligand-gated strand displacement mechanism for ZTP riboswitch transcription control.
E. J. Strobel*, L. Cheng, K. E. Berman, P. D. Carlson, J. B. Lucks*. Nature Chemical Biology. (2019).
Links: Journal, BioRxiv
4. Cotranscriptional folding of a riboswitch at nucleotide resolution.
K. E. Watters#, E. J. Strobel#, A. M. Yu, J. T. Lis, J. B. Lucks. Nature Structure and Molecular Biology. (2016). # = Equal contribution
Links: Journal, PDF