Recently we have begun to translate our fundamental discoveries into synthetic biotechnologies that improve global environmental and human health. We are particularly interested in harnessing the abilities of cells to monitor their environments into technologies that allow us to monitor our own internal and external environments. We have made dramatic progress in addressing this question by utilizing advances in “cell-free” synthetic biology to study synthetic RNA systems. In this approach, we use bacterial cell extracts to power gene expression reactions. By configuring biosensing mechanisms to regulate the expression of reporter genes (such as a fluorescent RNA or protein), these cell-free biosensors produce visual outputs when specific compounds are present. These systems can be freeze-dried and rehydrated upon use, allowing them to be stored and transported without cold chain, creating the possibility of an entirely new diagnostics platform based on cell-free synthetic biosensor reactions that is low-cost (<$1/test) and rapid (minutes).
Our early work in this area has focused on addressing the emerging water crisis that affects global public health. This includes using the fluoride riboswitch to develop a fluoride water quality diagnostic, developing a new concept of “metabolic biosensing” by which metabolism is used to break down a target molecule into one that can be sensed by a transcription factor, to create a diagnostic for the endocrine-disrupting herbicide atrazine, and devising a new platform that uses RNA genetic circuitry, called RNA Output Sensors Activated by Ligand INDuction (ROSALIND) to fix limitations of protein transcription factor specificity and sensitivity to create a panel of sensors for toxic metals (lead, copper, cadmium), antibiotics (tetracyclines, macrolides) and consumer products (salicylate, benzalkonium chloride). We have also begun to transition this technology out of the lab and into the world by demonstrating that these systems work in the field. Our current questions in this space are bold and ambitious: Can we sense anything, anywhere? We are addressing this through developing new sensors for human and environmental health, and new downstream signal processing components to make sensors more robust. We have also adapted this research area into developing new COVID-19 diagnostics that can scale to meet demand.
Featured Publications:
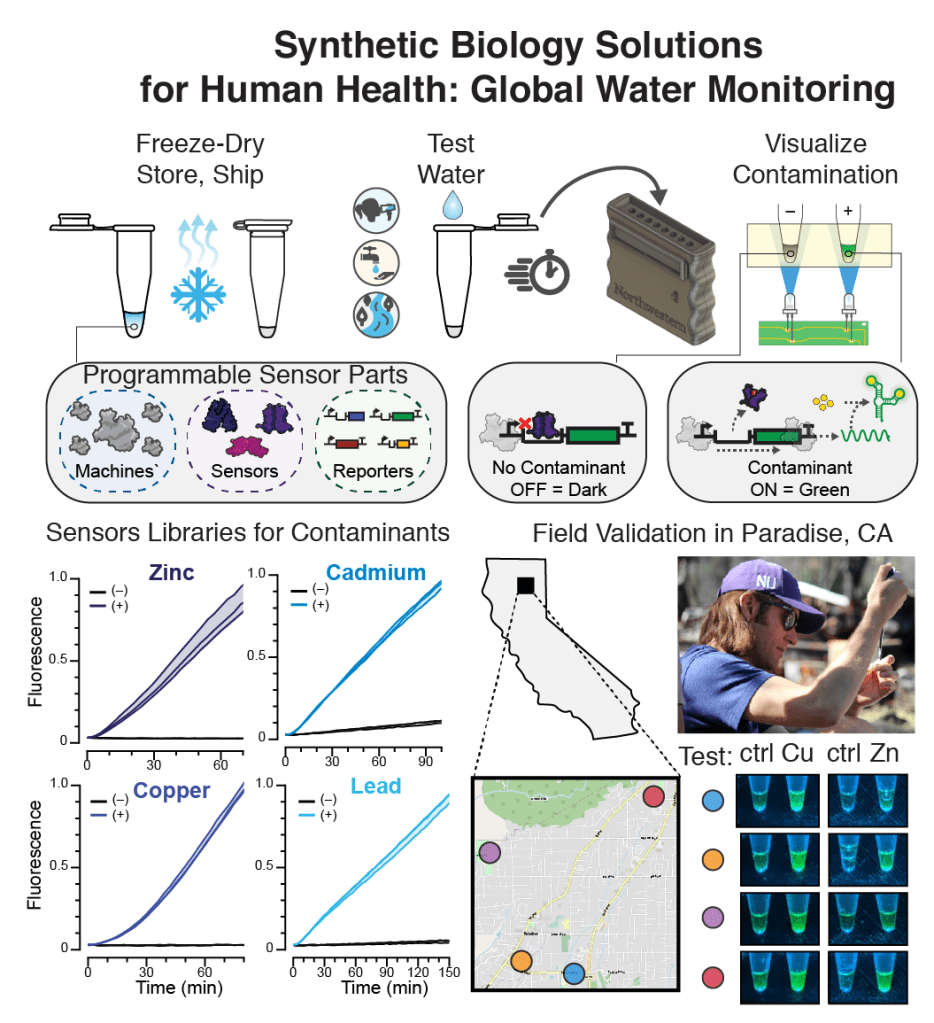
1. Rapid, low-cost detection of water contaminants using regulated in vitro transcription.
K. K. Alam#, J. K. Jung#, M. S. Verosloff, P. R. Clauer, JW. Lee, D. A. Capdevila, P. A. Pasten, D. P. Giedroc, J. J. Collins, J. B. Lucks*. (2019) # = Equal contribution
Links: BioRxiv
2. Point-of-use detection of environmental flouride via a cell-free riboswitch-based biosensor.
W. Thavarajah, A. D. Silverman, M. S. Verosloff, N. Kelley-Loughnane, M. C. Jewett, J. B. Lucks*. ACS Synthetic Biology.(2020)
Links: Journal, BioRxiv, PDF
3. Design and optimization of a cell-free atrazine biosensor.
A.D. Silverman, U. Akova, K.K. Alam, M.C. Jewett*, J. B. Lucks*. ACS Synthetic Biology.(2020)
Links: Journal, BioRxiv, PDF
4. PLANT-Dx: A molecular diagnostic for point of use detection of plant pathogens.
M. Verosloff, J. Chappell, K. L. Perry, J. R. Thompson, J. B. Lucks*. ACS Synthetic Biology. (2019). doi: 10.1021/acssynbio.8b00526
Links: Journal, PDF, BioRxiv