Our paper on Generating effective models and parameters for RNA genetic circuits is out in ACS Synthetic Biology. In this paper, Chelsea developed a simplified ODE model to model the dynamics of RNA genetic circuits. These ‘effective’ models essentially course grain a lot of the biochemical details of gene expression into simple mass-action kinetics that describe the synthesis and degradation of the relevant RNAs and proteins that make up a genetic network. Such models have been used for a long time for protein-based networks, though not much work showed that they could be used for RNA circuits. Chelsea’s first task was to do just that, and surprisingly they work very well after adding a few little details that are special to the RNAs we use to make the circuits. This is great news since the simpler the equations we can use, the more intuition we can gain about further engineering of these networks.
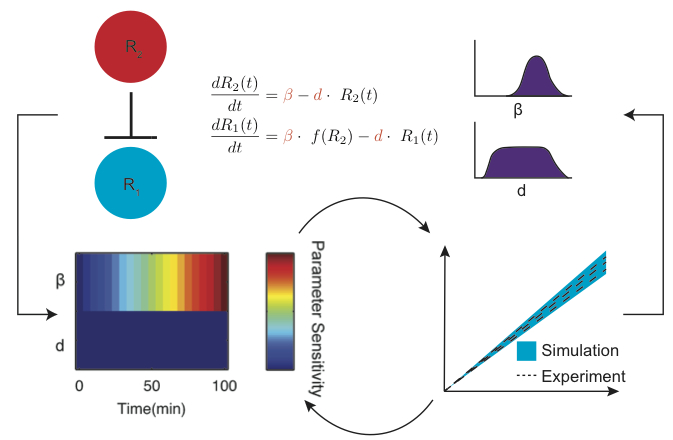
Even though these models worked well qualitatively, that is not quite the level of quantitative detail we want for the future when we think about applying these models to forward design RNA circuitry. So the next step was to find a way to parameterize the models. Unfortunately, since RNA-only genetic circuits are still pretty new (and not found in nature), there are not a lot of parameters to draw from. So we teamed up with Jeff Varner (Chelsea’s co-advisor) to apply methods from systems biology to help us design experiments that would parameterize the model. The method we endued up using uses sensitivity analysis on the structure of the ODEs to help us design experiments that can be used to find multiple parameters at once. In this way we could find all the parameters for our model in a limited number of experiments.
To perform the experiments, we turned to our new friend – cell free transcription-translation (TX-TL) reactions. TX-TL reactions mimic the cellular environment, but can be performed in a matter of hours. We ended up finding all the parameters in the model in four experiments that could be performed on a single plate in two hours using TX-TL reactions. What a good deal! We also went on to show that these experiments can actually tell you something about the TX-TL system you are using: by getting parameters from multiple batches of TX-TL you can see which parameters change, and it turns out that we think these changes have to do with the concentrations of the different molecules in the batches.
Overall we are very excited about this work. It’s our first step towards building a set of modeling tools that can be used to forward design RNA circuits. We also think the parameterization method is pretty powerful and could be used to parameterize all sorts of ‘parts’ for synthetic biology. To help everyone get started we put up our code on GitHub. Happy modeling/forking!