Research led by Lucks’ Lab PhD candidate Chelsea Hu was recently published in ACS Synthetic biology. The article, “Engineering a Functional Small RNA Negative Autoregulation Network with Model-Guided Design” expanded the repertoire of synthetic gene networks built from RNA regulators by constructing a transcriptional negative autoregulation (NAR) network out of small RNAs (sRNAs). NAR network motifs are core motifs of natural genetic networks, and are known for reducing network response time and steady state signal. In this work we use cell-free transcription–translation (TX-TL) reactions and a computational model to design and prototype sRNA NAR constructs. Using parameter sensitivity analysis, we design a simple set of experiments that allow us to accurately predict NAR function in TX-TL. We transfer successful network designs into Escherichia coli and show that our sRNA transcriptional network reduces both network response time and steady-state gene expression. This work broadens our ability to construct increasingly sophisticated RNA genetic networks with predictable function.
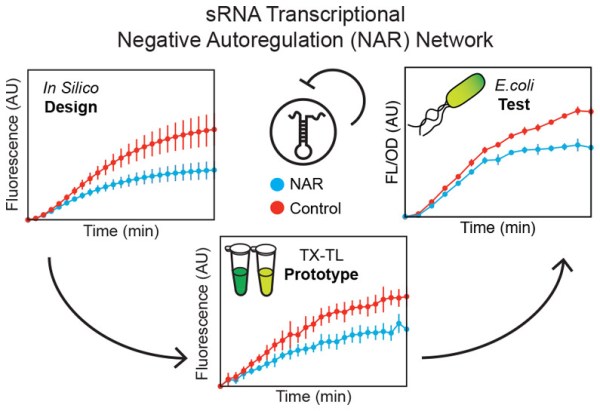
This work represents the first transcriptional sRNA negative autoregulatory circuit that functions both in cell-free reactions and in living cells. We anticipate this design-build-prototype-test network building process we used in this work will speed up network development cycles as the field moves toward more sophisticated synthetic network engineering.
Congratulations Chelsea!