We were recently invited to write a review on High-Throughput RNA Structure Determination by Nature Review Genetics and it just came out – congratulations to Eric and Angela! This was a great opportunity to revisit all of the powerful innovations that have happened in high throughput RNA structure determination throughout the years. For those new to the concept, the basic premise of the technique is to change the way measure RNA structures – they basically sacrifice resolution in favor of throughput and experimental flexibility. To do this they use biochemical reactions that react with RNAs in a structure-dependent fashion (see the figure) – these reactions covalently modify or cleave the RNA preferentially in regions that are un-paired or otherwise flexible. High throughput sequencing can then be used to map the locations of the modifications, creating a pipeline for measuring thousands of RNAs in a single experiment!
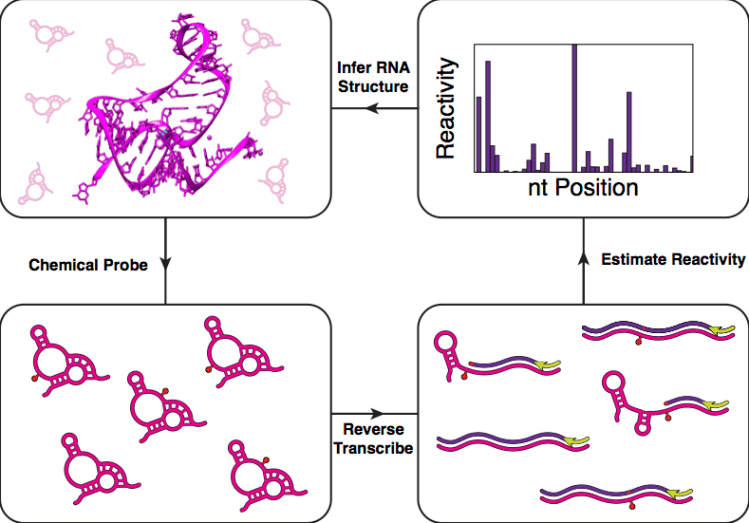
Figure: High throughput RNA structure determination begins by chemically or enzymatically probing RNA structures. Positions that are flexible or unpaired are modified by the probe or enzyme. Reverse transcription and sequencing are then used to map the probe positions and estimate a ‘reactivity’ for each nucleotide in the RNA. High reactivities indicate unstructured positions, while low reactivities could be structured, stacked, or interacting with other molecules. This information can be obtained for thousands of RNAs in a single experiment and powerfully be used to uncover structure-function relationships.
The throughput and flexibility of the approaches have really come a long way throughout the years. (In fact – Figure 1 of the review has a timeline of the history of this technology, stretching all the way back to the 1965 work of Robert Holley and team at Cornell that garnered a Nobel Prize for the interpretation of the genetic code!) Now these techniques have been used to map the structure of the ribosome, uncover high resolution structures of RNA molecules, and even map the structures of entire transcriptomes of cells! Our own recent developments of the SHAPE-Seq technology have allowed us to map how RNAs dynamically fold as they are being synthesized by RNA polymerase. With all of the rapid advances it is clear that these techniques are uncovering new layers of the RNA structure-function relationship in unprecedented ways.
With all those advances though it was high time to have a review that went through the details of the techniques to explain all of the nuances to new users. We took the opportunity to go back and read almost all of the literature in this area and summarize the available techniques by putting everything on the same conceptual footing (see the figure). We also learned a lot ourselves, and used the opportunity to take a deeper look into the meaning of the reactivity data generated by these techniques, and think about how the field can (and should!) come together to combine the best of all the techniques out there to come up with a really powerful experiment that is accessible to all biologists.
This is an exciting time for RNA biology. We can now use high throughput RNA structure probing techniques to ask questions once unimaginable. We are looking forward to being a part of the next wave of innovations in this exciting field! As always, we love to answer questions about how these techniques might be applicable to your problems. Please send us a note if you are interested!
New to High Throughput RNA Structure Determination? Check out this NIH video!